NANO-scale VIsualization to understand Bacterial virulence and invasiveness - based on fluorescence NANOscopy and VIBrational microscopy
|
| ||||
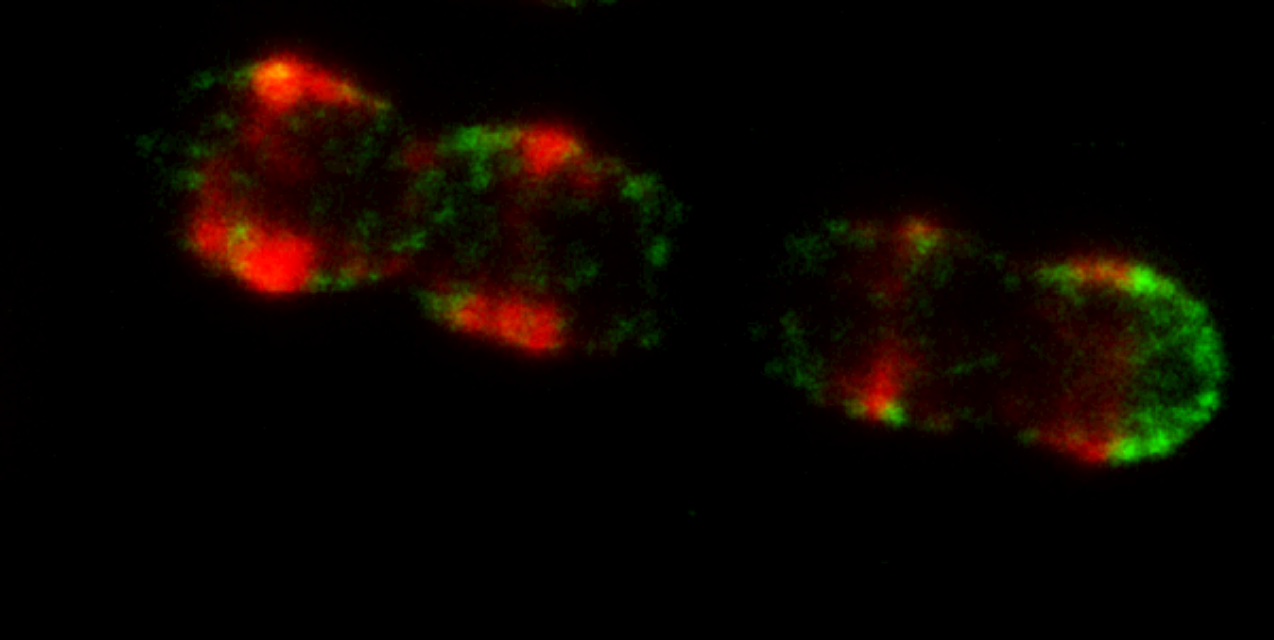
Background and motivationIn the biomedical field, fluorescence is the by far most widely used modality for cellular imaging, offering a unique combination of spatial and temporal resolution, sensitivity and specificity. There has been a remarkable development of fluorescence-based super-resolution microscopy techniques in the last decade. "Super-resolution" means that the resolution of the imaging is not limited by the wavelength of the light used to image a sample, and the main scientists behind this development were awarded the Nobel Prize in 2014. By the development of fluorescence-based super-resolution microscopy it has become possible to image cellular proteins marked with fluorescence emitters (fluorophores), with a ten-fold higher resolution than with any other fluorescence microscopy technique. In a previous EU FP7 project (Fluodiamon), this was used to analyze how certain proteins were spatially distributed in breast and prostate cancer cells, compared to in corresponding non-cancer cells, and was demonstrated as a new basis for cancer diagnosis. Two of the NanoVIB partners (KTH and KI) have also more recently applied super-resolution microscopy to study bacterial surface proteins of Streptococcus Pneumoniae (pneumococci). Pneumococcal infections represent a major cause of morbidity and mortality world-wide. These super-resolution microscopy studies could shed new light on how specific surface proteins of these bacteria are spatially distributed on the cells, and provided important evidence that the virulence (capacity to generate disease) and invasiveness of these bacteria is strongly coupled to such spatial protein distribution patterns (Figure 1). Clearly, imaging of these patterns with yet higher spatial resolution can lead to a significantly increased understanding of the mechanisms underlying pneumococcal virulence and invasiveness.
Now, MINFLUX, a next generation super-resolution concept, has recently been demonstrated, offering another order of magnitude higher spatial resolution than current super-resolution microscopy techniques (Figure 2). This means, spatial distribution patterns of proteins, which could be resolved within spatial scales of hundreds of nanometers by regular state-of-the-art fluorescence microscopes, and within tens of nanometers with super-resolution microscopy techniques, can now be resolved within a few nanometers, i.e. at a spatial scale corresponding to the size of the proteins themselves. Two of the NanoVIB partners (Abberior Instruments and IFNANO) have been closely involved in this development, originating from the 2014 Nobel Prize laureate Stefan W Hell and his research group in Göttingen. 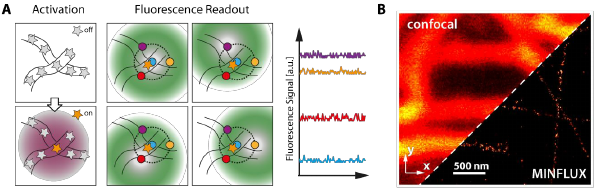
MINFLUX imaging can be performed with much lower irradiation doses and based on less bright fluorophores. This means that fluorophores in the near-infrared (NIR) spectral range (in our case 700-900nm) can now be used, which because of low brightness and photostability have not been eligible for super-resolution microscopy before. Imaging in the near infrared (NIR) can offer several distinct advantages: i) strongly reduced background scattering and autofluorescence from the sample, ii) lower phototoxicity, iii) deeper penetration depths, and iv) an additional spectral window, allowing for simultaneous imaging over multiple, spectrally separated color channels. To take MINFLUX into the NIR range however, requires detectors with high sensitivity in this spectral range. Such detectors will be developed in the NanoVIB project by the partner Pi Imaging, building further on their world-leading single photon counting detector technologies. To take full benefit of the information contained in the highly resolved protein localization patterns within cells, as obtainable by MINFLUX, we also want to overlay these patterns onto morphological images. To do this, without adding additional fluorophore markers that would sterically or spectrally interfere with the fluorophore reporters for MINFLUX, we will use in-elastic scattering from specific molecules already present in the cells. So-called stimulated Raman scattering (SRS) imaging requires sophisticated laser systems, with multiple, narrow linewidth emission lines, and precise tunability in the emission wavelength(s). The NanoVIB partner APE is in the absolute forefront of the development of such lasers. By further development and implementation of these laser systems into the MINFLUX prototype system, correlative imaging of nano-scale protein localization patterns and cellular morphology/chemical environment will be realized in the project. ObjectivesIn this coherent and interdisciplinary project, we will prototype a next-generation super-resolution microscope system based on the MINFLUX concept, capable to reveal intricate, detailed molecular mechanisms underlying inter- and intracellular processes and disease. As a lead application, we will demonstrate its unique capabilities to understand virulence and invasiveness of pathogenic bacteria, more precisely pneumococci. By concerted development of laser and detector technologies, microscopy devices and image acquisition procedures, we will be able to retrieve information, which is not within reach by any other microscopic or photonics-based technique. We will demonstrate how cellular nanoscale protein localization patterns can be resolved, which will not only help us revealing mechanisms of bacterial disease, but which also are likely to be of large relevance for many other diseases. Thus, the microscope system prototype will prove its broad applicability in biomedical research, as a tool to understand intra- and intercellular processes and the cellular origin of diseases. |
||||